
One of the major criticisms of scientific publishing is the long delay it takes between writing up the work and eventually getting it out in the public arena, with the fault shared variously between the authors, journal editorial boards, and peer reviewers. In the process, the data and results are usually embargoed. There is an excellent Nature news piece that described this last year. But it is invariably a pleasure to see the work out in print, and I am certainly glad that the multi-institutional work on understanding the huge Group B streptococcus (GBS) outbreak in Singapore in 2015 is now finally available (albeit behind a paywall) at the Clinical Infectious Diseases (CID) website.
To recap, unusually severe GBS infections were noticed in many local hospitals between March and July 2015. The majority of these cases were linked to the consumption of raw freshwater fish sold in porridge stalls in hawker centres, and were caused by a single strain of GBS, sequence type (ST) 283 – previously described to cause sporadic cases of severe human disease in Hong Kong, as well as disease in farmed fish in Asia. The outbreak ended soon after the public became aware of the link with consuming raw freshwater fish, and after our government agencies raised (and enforced) standards for selling fish for raw consumption.
What information is new in this academic publication?
- Firstly, there is a nice time series chart courtesy of the Saw Swee Hock School of Public Health team, showing a rise in blood isolates of GBS (signifying severe infection) from March 2015, whereas the number of GBS isolates cultured from all other body sites did not increase over the same time period.

A = GBS from blood. B = GBS from all other human sites. Data from most public sector hospital microbiology laboratories in Singapore. Snapshot from the CID publication.
- The corresponding clinical comparison between local adults infected with ST283 GBS (n=146) vs. non-ST283 GBS (non=262) in their blood showed that those infected with the outbreak strain of GBS:
- Were younger
- Were more likely to have consumed raw fish just prior to the infection
- Had fewer co-morbid conditions like diabetes mellitus or cancer
- Were more likely to have brain involvement (19.9% vs. 0%) and joint involvement (30.1% vs. 5.0%) but less likely to have skin (18.5% vs. 42.0%) or urinary tract (3.4% vs 11.8%) involvement.
- But were not more likely to die as a result of the ST283 GBS infection.
- We were also able to get ST283 GBS isolates from Hong Kong courtesy of Prof Margaret Ip, who studied the original cases in Hong Kong in 2006, and also from Bangkok courtesy of Prof Anucha Apisarnthanarak, who described a similar outbreak in Thailand in 2015.
- Excellent work by Dr Swaine Chen (GIS) and Prof Matthew Holden (St Andrews University, Scotland) analysing the entire genomes of the Singapore, Hong Kong and Thailand isolates showed that they were strikingly similar, and in fact, the Singaporean GBS isolates from humans (and the rare fish ST283 GBS we were able to obtain and sequence from Singapore General Hospital and the Environmental Health Institute) were clustered so closely together and with so few single nucleotide polymorphisms (SNPs) that the genomic data and results virtually suggested a point source outbreak in the Singapore context. The human and fish genomic results also virtually resolves the link between raw fish consumption and human disease.
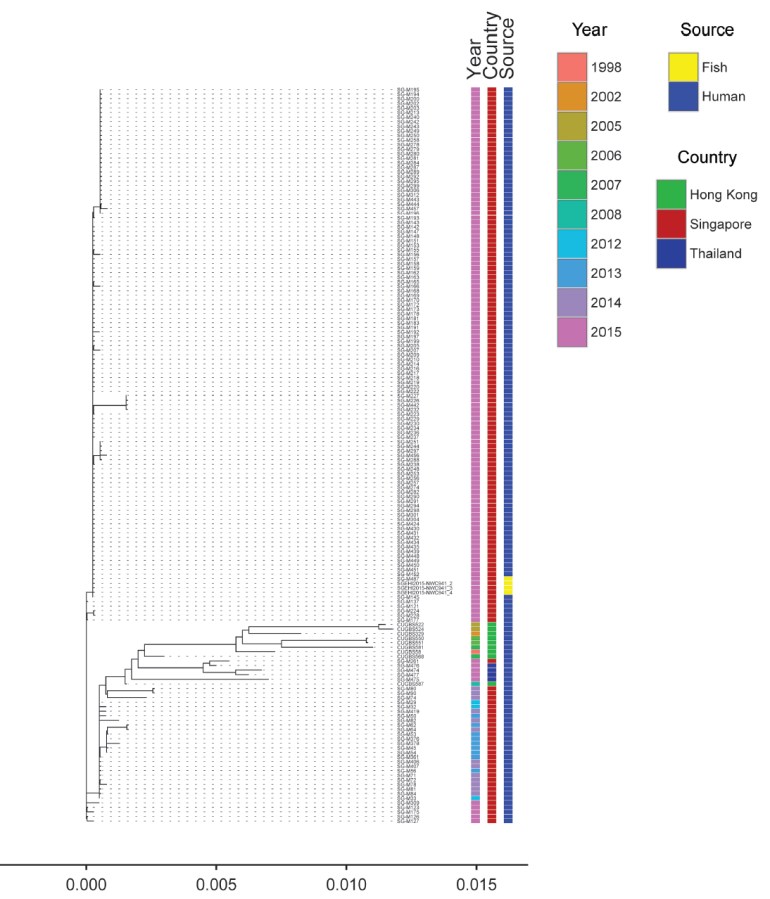
Phylogenetic tree from whole genome sequencing of GBS isolates obtained from fish (Singapore), and humans (Singapore, Hong Kong and Thailand). Snapshot from the CID publication.
- Although the outbreak occurred in 2015, ST283 GBS has already been causing human infections in Singapore since at least 2012, with increasing numbers each year until the fateful events of 2015.
What do we still not know after doing this work?
- The exact source of the outbreak remains unknown. At which point were the fish infected (at the farms themselves, or somewhere during the logistics chain down to Singapore)?
- Why is the clinical presentation different between this “special” clone of GBS as compared to other GBS infections?
- We know lots of people (used to) consume raw yusheng at porridge stalls – far more than the number actually infected with ST283 GBS. Why did the rest not get the infection? Was it a matter of lower infectious “dose” or some kind of immune defect? This is also not clear at present.
[…] still a significant number of unanswered questions) has just been published, as described in an earlier blog post. The almost 2-year delay between the outbreak and publication of the work is disappointing, and […]
LikeLike